Carbon monooxide
In this example, how to draw molecular orbital(s) is explained by taking a carbon monooxide (CO) molecule as an example.
Structural optimization
To start off, we first optimize the structure by using the quenched molecular dynamics (QMD) using the following input file (nfinp_opt):
WF_OPT DAV
GEO_OPT QMD
NTYP 2
NATM 2
GMAX 6.00
GMAXP 20.00
MIX_ALPHA 0.7
WIDTH 0.0010
EDELTA 1.D-09
NEG 8
FMAX 1.D-3
CELL 15.00000000 15.00000000 15.00000000 90.00000000 90.00000000 90.00000000
&ATOMIC_SPECIES
C 12.011 pot.C_pbe1
O 15.999 pot.O_pbe1
&END
&ATOMIC_COORDINATES CARTESIAN
0.000000000000 0.000000000000 0.000000000000 1 1 1
2.200000000000 0.000000000000 0.000000000000 1 1 2
&END
Run STATE and confirm that the structural optimization is successfully finished by inspecting the input file (nfuot_opt) by:
$ grep -A1 f_max nfout_opt
Then you may get the following:
NIT TotalEnergy f_max f_rms edel vdel fdel
1 -22.28228624 0.032705 0.032674 0.13D-09 0.24D-07 0.13D-09
--
NIT TotalEnergy f_max f_rms edel vdel fdel
2 -22.28247708 0.025505 0.025471 0.16D-10 0.46D-07 0.16D-10
--
NIT TotalEnergy f_max f_rms edel vdel fdel
3 -22.28269805 0.012213 0.012175 0.64D-10 0.57D-07 0.64D-10
--
NIT TotalEnergy f_max f_rms edel vdel fdel
4 -22.28275321 0.004811 0.004776 0.61D-10 0.47D-07 0.61D-10
--
NIT TotalEnergy f_max f_rms edel vdel fdel
5 -22.28275321 0.004810 0.004770 0.61D-10 0.24D-07 0.61D-10
--
NIT TotalEnergy f_max f_rms edel vdel fdel
6 -22.28275706 0.003622 0.003578 0.33D-09 0.75D-07 0.33D-09
--
NIT TotalEnergy f_max f_rms edel vdel fdel
7 -22.28276111 0.001571 0.001536 0.91D-11 0.35D-07 0.91D-11
--
NIT TotalEnergy f_max f_rms edel vdel fdel
8 -22.28276145 0.000925 0.000888 0.11D-09 0.28D-07 0.11D-09
We then use the geom2nfinp to generate a new input file for the following up calculation with the optimized geometry as:
$ geom2nfinp -i nfinp_opt -g GEOMETRY -o nfinp_prtwfc
Make sure that the GEOMETRY file exsit in the working directory.
Wave function plot
We then modify the input file nfinp_prtwfc as:
TASK PRTWFC
WF_OPT DAV
NTYP 2
NATM 2
GMAX 6.00
GMAXP 20.00
MIX_ALPHA 0.7
WIDTH 0.0010
EDELTA 1.D-09
NEG 8
CELL 15.00000000 15.00000000 15.00000000 90.00000000 90.00000000 90.00000000
&ATOMIC_SPECIES
C 12.011 pot.C_pbe1
O 15.999 pot.O_pbe1
&END
&ATOMIC_COORDINATES CARTESIAN
0.015722761422 0.000000149539 -0.000000142524 1 1 1
2.188120977445 0.000000097882 -0.000000268744 1 1 2
&END
&PLOT
IK 1
IBS 5
IBE 7
FORMAT XSF
&END
You can see the the atomic coordinates are different from the original ones. In addition you can find the line:
TASK PRTWFC
to tell the code to plot the wave function in real space and the block:
&PLOT
IK 1
IBS 5
IBE 7
FORMAT XSF
&END
to tell which orbitals to be plotted. In this case, we are going to plot 5th to 7th wave functions, which correspond to highest occupied molecular orbital (HOMO) and doubly degenerate lowest unoccupied orbitals (LUMOs) and the output format is XSF.
By executing STATE by using the above input file (nfinp_prtwfc), you may obtain the following wave functions:
nfwfn_kpt0001_band0005_re.xsf
nfwfn_kpt0001_band0005_im.xsf
nfwfn_kpt0001_band0006_re.xsf
nfwfn_kpt0001_band0006_im.xsf
nfwfn_kpt0001_band0007_re.xsf
nfwfn_kpt0001_band0007_im.xsf
The naming convention is:
nfwfn_kpt[kpoint_index]_band[band_index]_[re/im].xsf
where “re” (“im”) indicates that it is the real (imaginary) part of the wave function.
By using VESTA the real part of the wave functions for HOMO and LUMO can be visualized as:
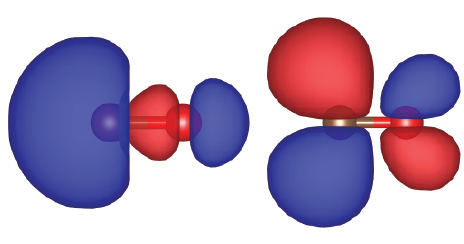
An energy diagram of CO can be found in the Supplementary Material of Wella et al. J. Chem. Phys. 152, 104707 (2020).